Introduction
GoalThe aim is to : get familiar with datasets produced by Next Generation Sequencing (NGS).
- accessing data in GEO / SRA/ ENA databanks from an accession number in a publication
- searching for public datasets in GEO
- understanding the FASTQ format used for reads files
- Assess the overall quality of a FASTQ read file
For this training, we will use various ChIP-seq datasets. ChIP-seq is a functional genomics assay to find all the regions in a genome bound by a particular protein (e.g. a transcription factor).
- Bacteria: ChIP-seq dataset produced by Myers et al [PDF] involved in the regulation of gene expression under anaerobic conditions in bacteria. We will focus on one factor: FNR.
- Zebrafish: ChIP-seq produced by Paik et al [PDF] to study the factor Cdx4 during development. We will focus on H3K27ac in bud-stage embryo.
Download a dataset
1 - Obtaining an identifier for a chosen dataset
NGS datasets are (usually) made freely accessible for other scientists, by depositing these datasets into specialized databanks. Sequence Read Archive (SRA) located in USA hosted by NCBI, and its european equivalent European Nucleotide Archive located in England hosted by EBI both contains raw reads.
Functional genomic datasets (transcriptomics, genome-wide binding such as ChIP-seq,...) are deposited in the databases Gene Expression Omnibus (GEO) or its European equivalent ArrayExpress.
Within an article of interest, search for a sentence mentioning the deposition of the data in a database (tip: automatically search in the pdf "GEO" or "SRA"). In the Bacteria dataset [PDF] , the following sentence can be found at the end of the Materials and Methods section:
"All genome-wide data from this publication have been deposited in NCBI’s Gene Expression Omnibus (GSE41195)."
We will thus use the GSE41195 identifier to retrieve the dataset from the NCBI GEO (Gene Expression Omnibus) database.
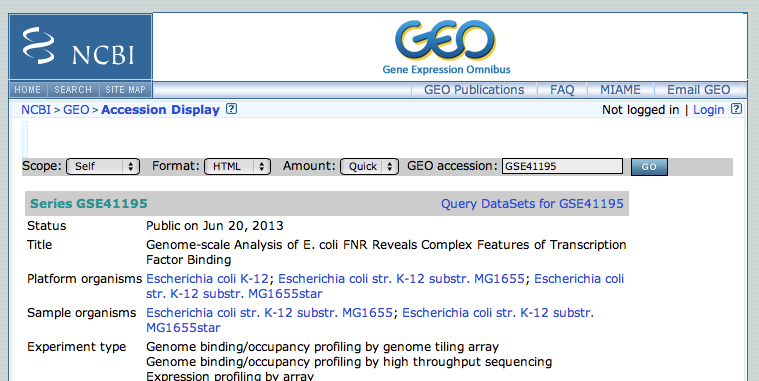
2 - Accessing GSE41195 from GEO
- Search in google GSE41195. Click on the first link to directly access the correct page on the GEO database.
- This GEO entry is a mixture of expression analysis and chip-seq. At the bottom of the page, click on the subseries related to the chip-seq datasets. (this subseries has its own identifier:GSE41187)
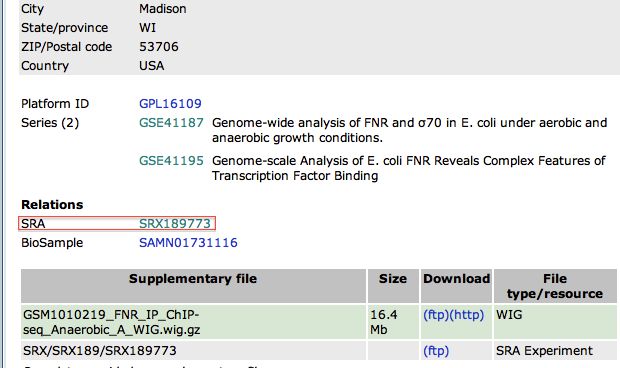
- From this page, we will focus on the experiment FNR IP ChIP-seq Anaerobic A. At the bottom of the page, click on the link "GSM1010219 - FNR IP ChIP-seq Anaerobic A"
- In the new page, go to the bottom to find the SRA identifier. This is the identifier of the raw dataset stored in the SRA database.
- Copy the identifier SRX189773 (do not click on the link)

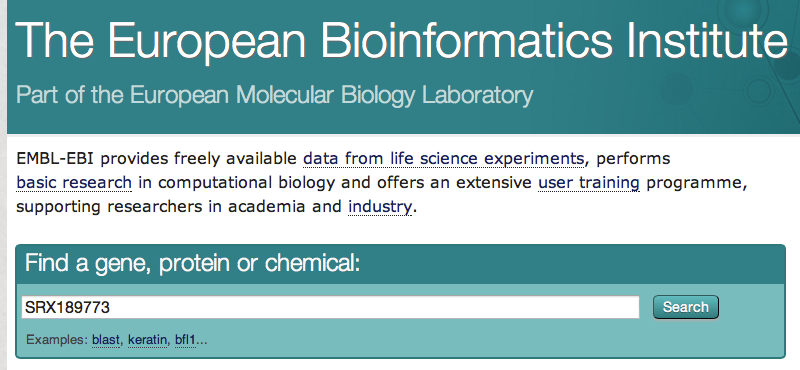
3 - Downloading FASTQ file from the EBI database
- Go to the EBI website. Paste your identifier (SRX189773) and click on the button "search"
- Click on the first result. On the next page, there is a link to the FASTQ file. Download this file (= save this file on your computer). As FASTQ files are large, this will be the only sequence file that we download for this course.
- For the second article, redo all the steps above and find the link to the FASTQ file (without downloading it...)
Searching GEO
1 - Searching the GEO
- Go to the NCBI website and select GEO datasets as the database to search.
- Search the term chip-seq
- The left part of the page aims as enabling filters on the current search. It's possible to filter on the organism, study type, tissue... Filter the results to obtain only zebrafish datasets.
As of today, 11702 datasets in GEO match the term "chip-seq". This number keeps growing...
Under organism, click on the link Select.... By searching zebrafish, the system automatically proposes "Danio rerio", the latin name of this organism. This filter limits the search to 46 datasets.
2 - Analyze a GEO entry
- Click on the dataset "Zic3 interacts with distant regulatory elements to regulate zebrafish developmental genes".
What was the overall design ?
Is the dataset published (see Citation section) ?
On which sequencing platform was the data produced ?
On this example in zebrafish, the data was publicly released on Feb 27, 2014. This dataset is related to a publication in the journal Plos Genetics. The overall design is ChIP-seq targeting the protein Zic3 in wild type and sqet33 transgenic. Sequencing was performed on a Illumina Genome Analyzer, in Singapore.
3 - Search the GEO by keyword
- Go back to the search page, enter the term hox* and filter by organism zebrafish
- Go back to the search page, enter the term CLIP-Seq
44 datasets are related to one of the Hox gene family in zebrafish
Look at the top right of the page. 6 datasets related to CLIP-seq are counted.
Viewing raw (FASTQ) data
1 - Obtain a FASTQ file
FASTQ files are generally larger than 1 Go. They cannot be opened by usual programs (and you should avoid trying opening them, as it could really freeze your computer). To view such big text files - text as in not binary files - you can use the terminal.
You should already have downloaded the Bacteria dataset FASTQ file on your computer. Do you remember where ?
If you haven't downloaded it yet, you can obtain it from the ENA database with accession SRX189773.
Save it on your Desktop.
2 - Viewing the FASTQ file
- Go to the Desktop folder
cd Desktop
- List the content of this folder. You should see the downloaded file SRR576933.fastq.gz. How large is the file ?
ls -lth -rw-r--r-- 1 mthomas csb 121M avril 1 21:45 SRR576933.fastq.gz
- The file is zipped. Unzip the file
gunzip SRR576933.fastq.gz
- Check again the content of the folder How large is the fastq file ?
ls -lth -rw-r--r-- 1 mthomas csb 453M avril 1 21:45 SRR576933.fastq
- View the 20 first lines (means the 5 first reads, as there are 4 lines per read)
head -n 20 SRR576933.fastq
3 - Understanding the FASTQ file
- Use google to search for the specifications of the FASTQ format. Your FASTQ file is encoded in Sanger FASTQ.
Quality control of the reads and statistics
1 - Getting the FASTQC report
Before you analyze the data, it is crucial to check the quality of the data. We will use the standard tool for checking the quality of data generated on the Illumina platform: FASTQC.
- First check the help page of the program to see its usage and parameters.
fastqc --help
- Run fastqc program on your dataset.
fastqc SRR576933.fastq.gz
- Check the files output by the program.
ls SRR576933_fastqc SRR576933_fastqc.zip SRR576933.fastq.gz
- Access the result.
cd SRR576933_fastqc ls fastqc_data.txt fastqc_report.html Icons Images summary.txt
- Open the HTML file fastqc_report.html in Firefox.
How many reads are present in the file ?
What is the read length ? Is the overall quality good ?
Are there any concerns raised by the report ? If so, can you tell where the problem might come from ?
There are 3 603 544 reads of 36bp. The overall quality is good, although it drops at the last position, which is usual with Illumina sequencing, so this feature is not raising hard concerns. There are several "red lights" in the report. In particular, the per sequence GC content and the duplication level are problematic. If you check the "overrepresented sequences", you'll notice a high percentage of adapters (29% !). Ideally, we would remove these adapters (=trim) the reads, and then re-run FASTQC. Results.
What are the adapters ?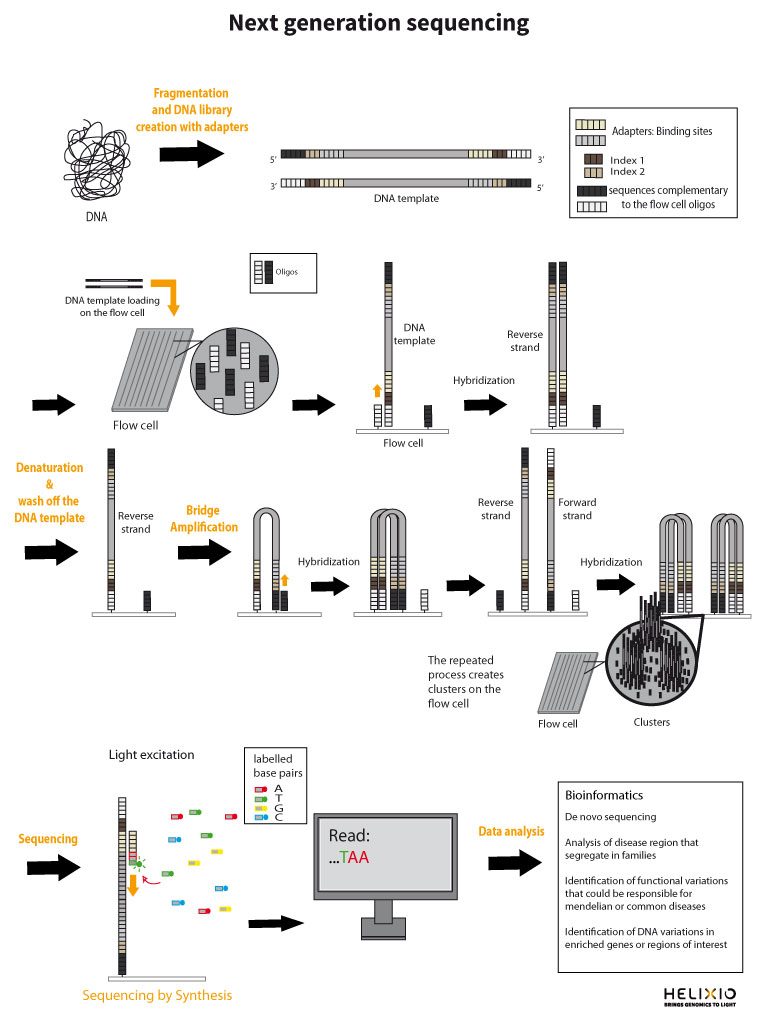
2 - Organism length
Knowing your organism size is important to evaluate if your dataset has sufficient coverage to continue your analyses. For the human genome (3 Gb), we usually aim at the very least 10 Million reads for ChIP-seq.
- Go to the NCBI Genome website, and search for the organism Escherichia coli
- Click on the Escherichia coli str. K-12 substr. MG1655 to access statistics on this genome.
Do both FASTQ files contain enough reads for a proper analysis ?
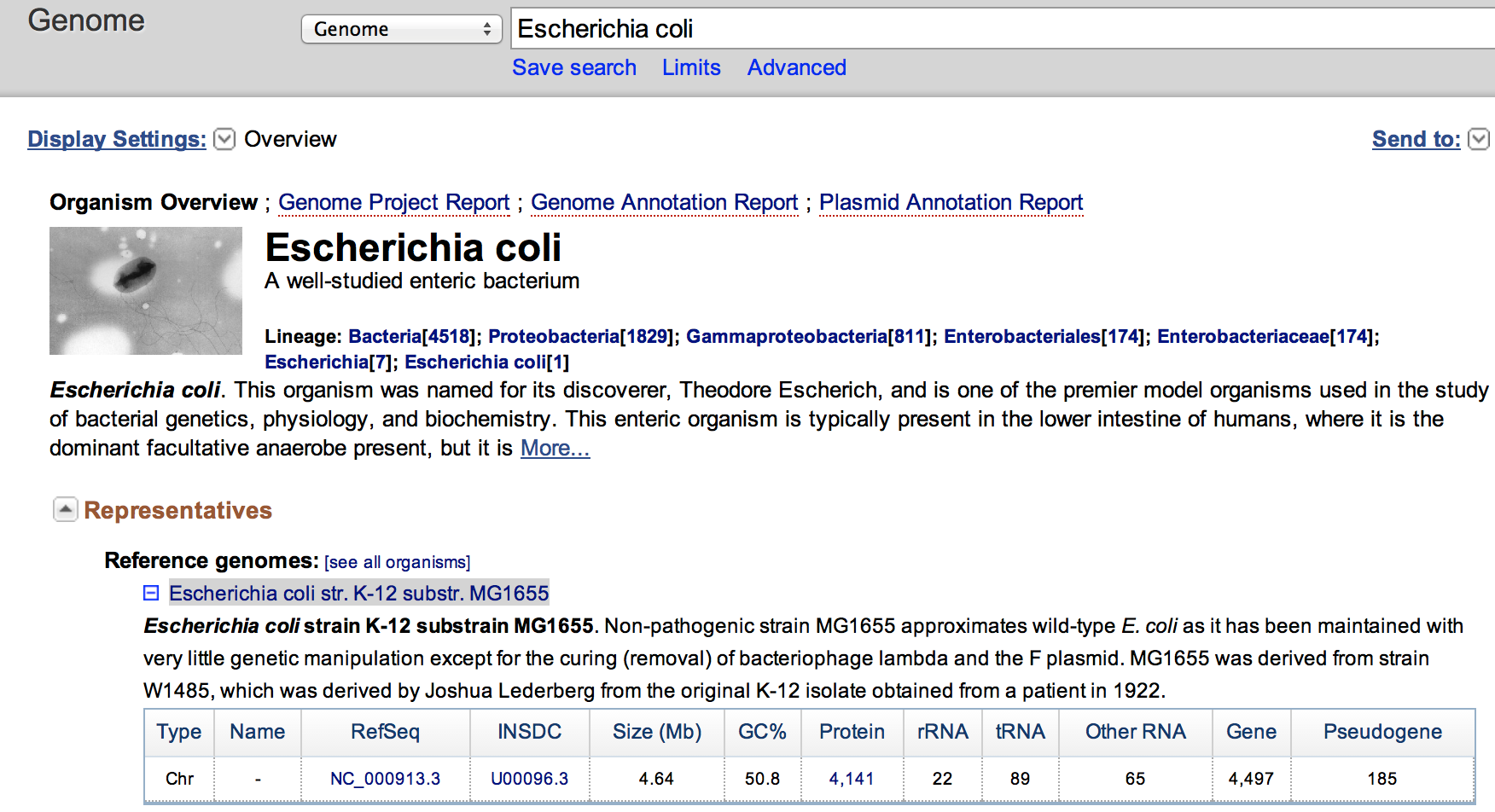
The genome is 4.64 Mbase. The file has 3.6 M reads. As we aim for 10 M for 3 Gb when working with human, the dataset here should enough reads for proper analysis.